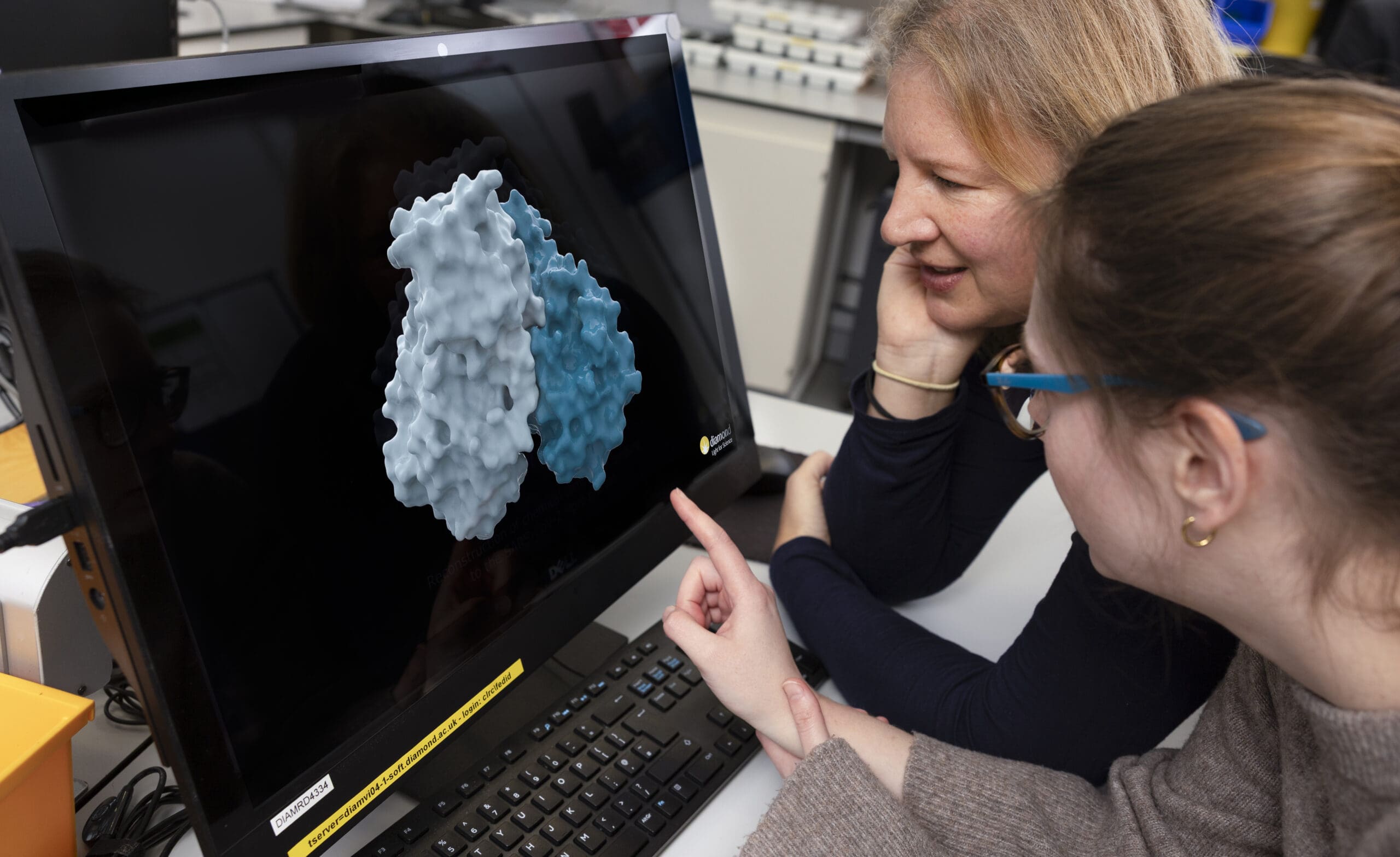
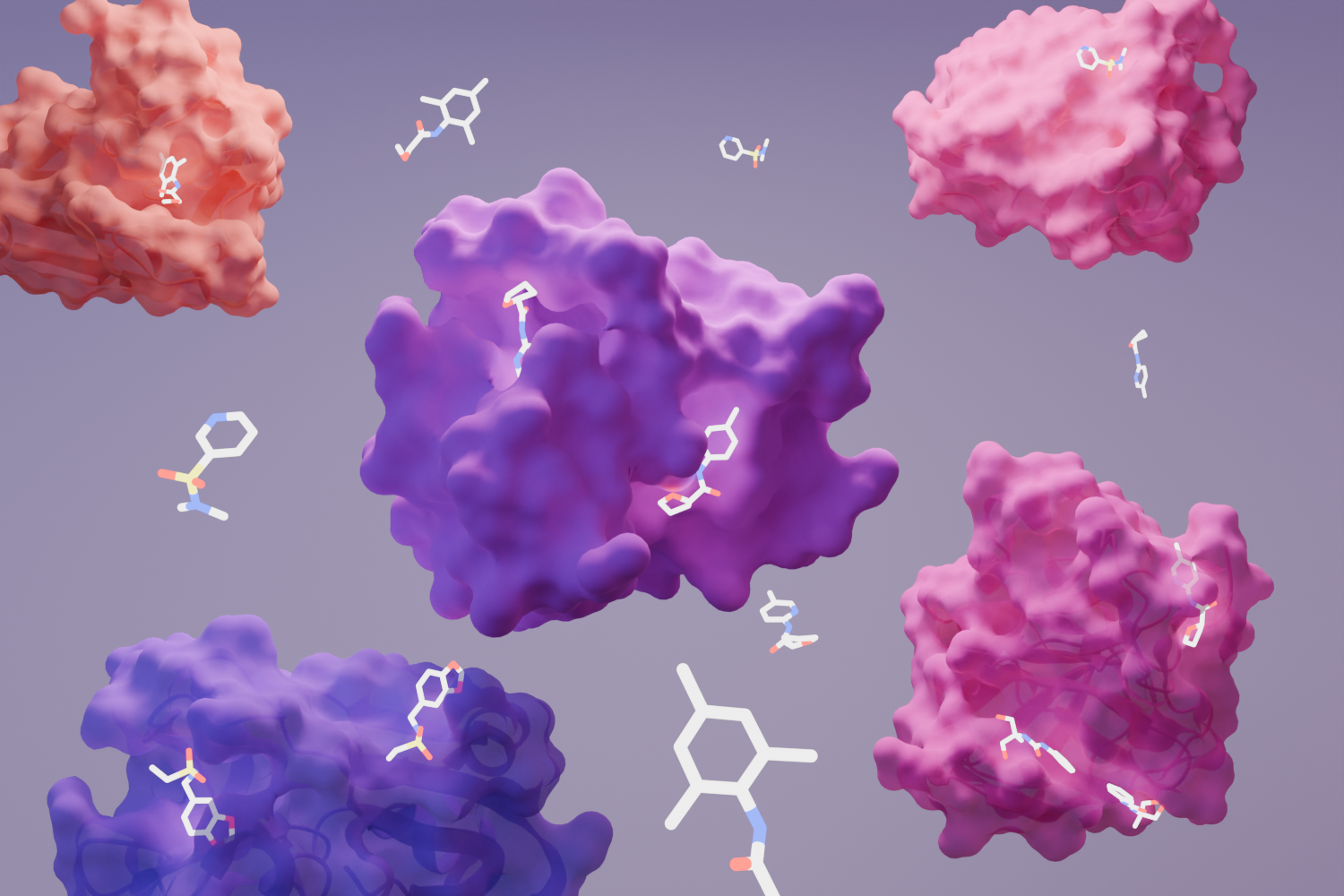
The UK‑led OpenBind initiative has reached a major milestone with the announcement of the release of its first publicly available dataset and predictive AI model, a groundbreaking step toward accelerating the discovery of new medicines using artificial intelligence.
The release showcases how engineering the production of AI-ready data is not only feasible but essential to evolving AI tools for scientific fields, which all suffer from a lack of data. With this OpenBind release, both high‑quality, standardised experimental data, and a newly trained predictive model, OpenBind v1, will become freely accessible to researchers worldwide, for immediate use in therapeutic discovery and to drive the next generation of AI models.

Credit: Stuart March-DNDi
While AI has introduced a step‑change in predictive accuracy for protein structures, its impact on drug discovery has remained muted, limited above all by the global shortage of reliable experimental data measuring in atomic detail how molecules of drug discovery bind to disease‑related proteins. OpenBind aims to fill this critical gap. Led by Diamond Light Source, the collaboration of structural biologists and AI specialists – supported in its foundation phase by the Department for Science, Innovation and Technology (DSIT) – is the first initiative to generate these essential datasets at industrial scale, openly and continuously, and designed specifically for AI.
This first release demonstrates that OpenBind’s pipeline is now operational, having generated 800 high-quality measurements in only 7 months – in the past, such large datasets took years to be produced and released. This integrated operation combines automated chemistry, robust binding measurements and high‑throughput crystallography at Diamond’s XChem Fragment Screening facility with an engineered data release process and AI model training using UK’s Isambard-AI compute cluster. It lays the groundwork for transformative progress in drug discovery, with future data tranches planned to address global‑health challenges such as COVID‑19, malaria, dengue, Zika, and cancer, where rapid development of new treatments remains vital.
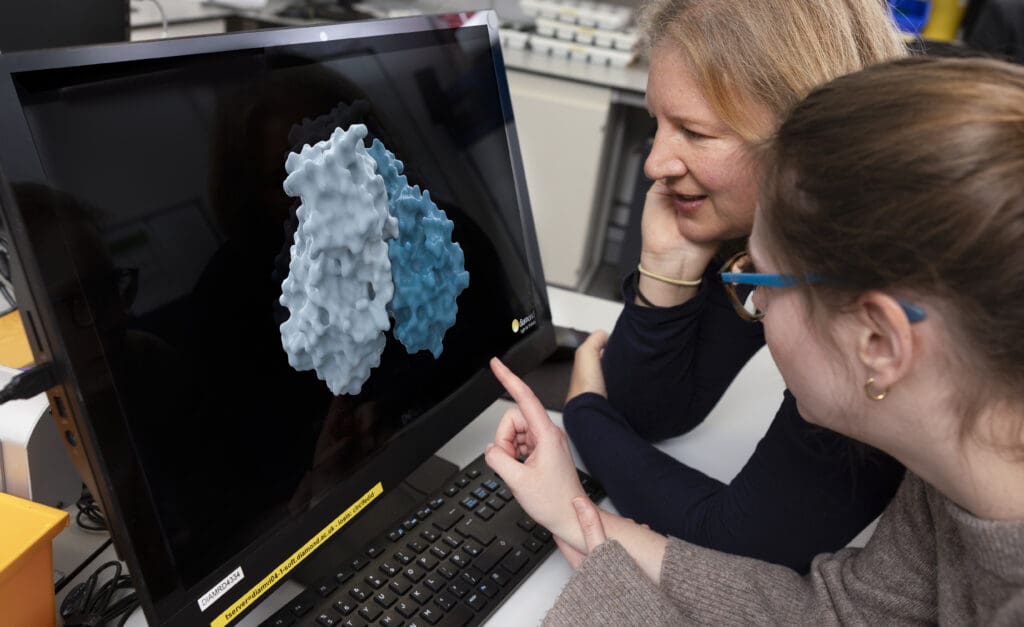
Credit: Stuart March – DNDi
“AlphaFold2 revolutionised protein structure prediction by leveraging decades of experimental data on protein structures in the PDB,” states Prof. Mohammed Alquraishi, Columbia University.“ The equivalent of such a dataset for protein-drug complexes does not yet exist, but OpenBind aims to create it, and in the process create the next generation of computational tools for modeling interactions between drugs and proteins.”
The initial dataset also reflects invaluable learning from the initiative’s early experimental cycles. Standardised workflows, strong metadata practices, and high levels of automation have proven crucial in ensuring the consistency and reproducibility required for AI, while highlighting opportunities to further streamline data handling and release frequency.
“High-quality experimental data is essential for developing new and improved AI models, and this first data release shows that OpenBind now has this foundation in place. We’re enabling AI to improve model performance and guide future experiments, helping to accelerate discovery,” says Dr. Fergus Imrie, University of Oxford. “The lessons from these early cycles are already helping us improve the speed, consistency, and reproducibility of the pipeline, which will be critical as OpenBind grows.”
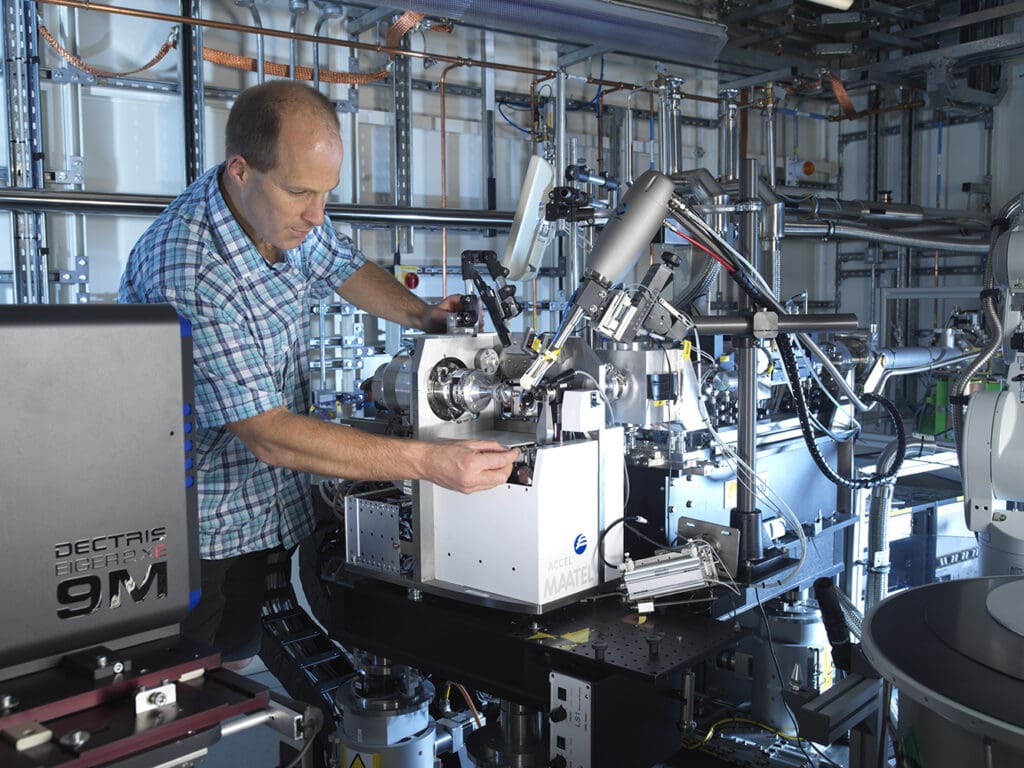

Credit: Stuart March – DNDi
“We couldn’t have made such rapid progress without the contributions of our consortium members and operational team,” says Prof. Frank von Delft, Principal Scientist at Diamond. “Their expertise and commitment have enabled us to reach this ambitious milestone. We will now implement the lessons from this foundation phase to ramp up a long-term operation that links high-volume production of AI data with active discovery projects.”
Building on this foundation, OpenBind will expand to include many more targets, larger chemical series, and deeper datasets, alongside community blind‑challenges that will validate AI models for newly generated experimental data. Ultimately, OpenBind aims to create a global, open data engine capable of supporting the development of faster, more accurate, and more equitable therapeutics.